Naftiko turns your existing data and APIs into governed capabilities for AI, giving you the control you need without throwing away prior investment. Few sectors stand to benefit more from this approach than healthcare, where decades of legacy integration work represent both an enormous asset and an ongoing challenge. Let’s explore how Naftiko helps modernize and standardize healthcare integration across Europe.
The Healthcare Integration Challenge
Naftiko is focused on standardizing and governing software integrations, and there are few sectors where this matters more than in healthcare. We want to explore how Naftiko can help enable the work at the European Health Data Space (EHDS), an EU initiative aimed at creating a unified framework for secure, cross-border access to patient health data, facilitating both personal care (primary use) and research and innovation (secondary use). By leveraging the Naftiko Capabilities specification and the Naftiko Framework, healthcare providers can transform legacy data, and then govern the integration of that data across systems and borders.
European hospitals run on HL7 v2. These pipe-delimited messages flow between EMR systems, national patient registries, and lab systems across the continent. The data is rich, the volume is massive, and the format is deeply embedded. Nobody is ripping it out. But the world is moving toward OpenEHR, a structured, archetype-based clinical data model that enables interoperability, FHIR bridging, and the kind of standardized data access the EHDS demands.
The question is not whether to modernize. The question is how to do it without losing what already works.
What Naftiko Capabilities Do
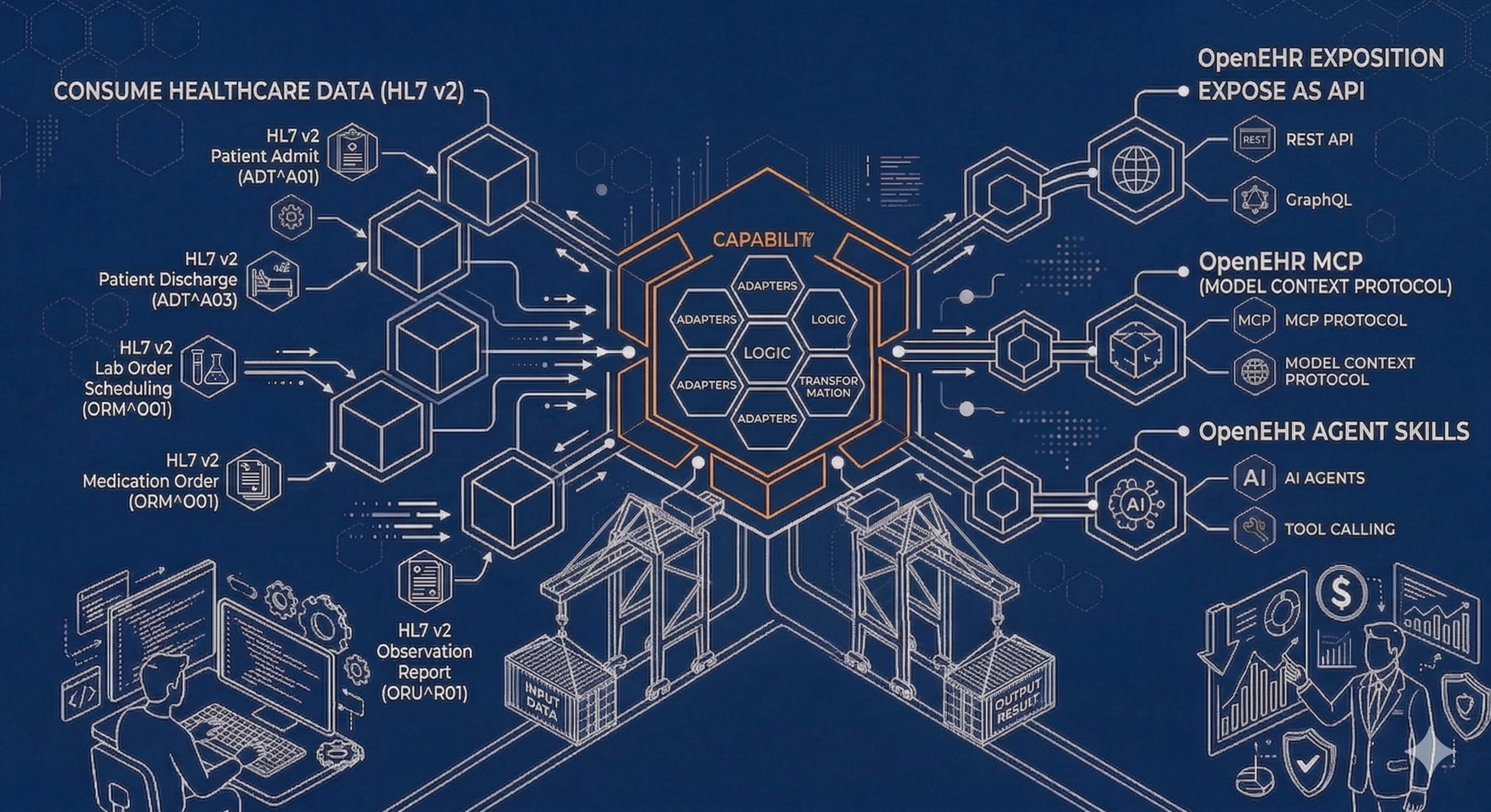
Naftiko aligns integrations with business outcomes using declarative, open-source, machine-readable and human-readable capabilities expressed as YAML. A Naftiko Capability consumes your existing data and APIs and produces what you need for AI integration. In this healthcare scenario, we use Naftiko Capabilities to consume classic HL7 v2 patient data and transform it into standardized, interoperable OpenEHR compositions.
Each capability declares three things:
- What it consumes – the existing HL7 v2 gateway and its message segments
- What it produces – the OpenEHR COMPOSITION posted to the clinical repository
- What it exposes – an MCP tool that an AI agent can discover and invoke
No custom code. No middleware. Just a YAML file that the Naftiko Framework executes in a Docker container.
Patient Admit: ADT^A01 to OpenEHR Encounter
The most fundamental hospital event is a patient admission. An ADT^A01 message fires when a patient is admitted, carrying identity, location, attending physician, and admitting diagnosis. Here is the Naftiko Capability that transforms it.
The Capability
naftiko: "1.0.0-alpha1"
info:
label: HL7 v2 Pipe ADT^A01 Patient Admit to OpenEHR
description: >-
Transforms raw pipe-delimited HL7 v2 ADT^A01 Patient Admit messages into
OpenEHR-compatible COMPOSITION resources. Consumes a hospital HL7 v2
MLLP gateway that returns native pipe-separated HL7 v2 messages (not JSON)
and produces an OpenEHR encounter composition with admin_entry for admission
details and problem_diagnosis for admitting diagnosis.
tags:
- hl7v2
- openehr
- adt-a01
- patient-admit
- healthcare
capability:
consumes:
- namespace: hl7-gateway
type: http
baseUri: "https://hl7-gateway.hospital.example.eu/api/v1"
description: >-
Hospital HL7 v2 message gateway. Returns raw pipe-delimited
HL7 v2 ADT^A01 Patient Admit messages as text/plain from the
integration engine.
authentication:
type: bearer
token: "{{HL7_GATEWAY_TOKEN}}"
resources:
- name: adt-events
path: "/adt/events"
operations:
- name: get-admit-event
method: GET
- namespace: openehr-repo
type: http
baseUri: "https://ehrbase.hospital.example.eu/rest/openehr/v1"
description: >-
OpenEHR clinical data repository (EHRbase). Receives transformed
COMPOSITION resources conforming to openEHR-EHR-COMPOSITION.encounter.v1.
authentication:
type: bearer
token: "{{EHRBASE_TOKEN}}"
resources:
- name: compositions
path: "/ehr/{{ehrId}}/composition"
operations:
- name: create-composition
method: POST
exposes:
- type: mcp
port: 3010
namespace: hl7-openehr-tools
description: >-
MCP tools for transforming HL7 v2 ADT^A01 Patient Admit messages
into OpenEHR encounter compositions.
tools:
- name: transform-adt-a01
description: >-
Fetches an ADT^A01 Patient Admit message from the HL7 gateway,
transforms it into an OpenEHR encounter composition with admission
details and diagnosis, and posts it to the OpenEHR repository.
inputParameters:
- name: messageId
type: string
required: true
description: "HL7 message control ID (MSH-10)"
- name: ehrId
type: string
required: true
description: "Target OpenEHR EHR identifier"
steps:
- name: fetch-admit
type: call
call: hl7-gateway.get-admit-event
with:
messageId: hl7-openehr-tools.messageId
- name: post-composition
type: call
call: openehr-repo.create-composition
with:
ehrId: hl7-openehr-tools.ehrId
Read it top to bottom: consume the HL7 gateway, consume the OpenEHR repository, expose an MCP tool that orchestrates the two. The steps section fetches the HL7 message, then posts the transformed composition. That is the entire integration.
What Gets Consumed: The HL7 v2 Message
This is what comes off the wire. A real ADT^A01 from a European hospital, with the patient’s national identifier, ward assignment, attending physician, and admitting diagnosis coded in ICD-10:
MSH|^~\&|EMR_SYSTEM|CENTRAL_HOSPITAL|NATIONAL_REGISTRY|HEALTH_AUTHORITY|20240315082300||ADT^A01^ADT_A01|MSG00001|P|2.5|||AL|NE||UTF-8
EVN|A01|20240315082300|||^Lindqvist^Anna^^^Dr.
PID|1||7812150423^^^NATIONAL_ID&2.16.840.1.113883.4.1&ISO~10045^^^EMR&LOCAL||Mueller^Erik^Johan||19781215|M|||Hauptstrasse 12^^Berlin^^10115^DE^H||+4930123456|||M
PD1|||City Medical Center^^12345|^Weber^Maria^^^Dr.
PV1|1|I|WARD-23^Room-4^Bed-2|E|||^Lindqvist^Anna^^^Dr.|^Bergman^Lars^^^Dr.||MED|||||||^Lindqvist^Anna^^^Dr.|INP|V201500123456||||||||||||||||||||||A|||20240315082300
DG1|1||J18.9^Pneumonia, unspecified^ICD10|Pneumonia|20240315|A
Every field matters. PID-3 carries the patient’s national identifier (7812150423) with the appropriate OID namespace. PV1-3 encodes the ward, room, and bed (WARD-23^Room-4^Bed-2). DG1-3 carries the ICD-10 diagnosis code (J18.9 – unspecified pneumonia). This is the reality of healthcare data in production.
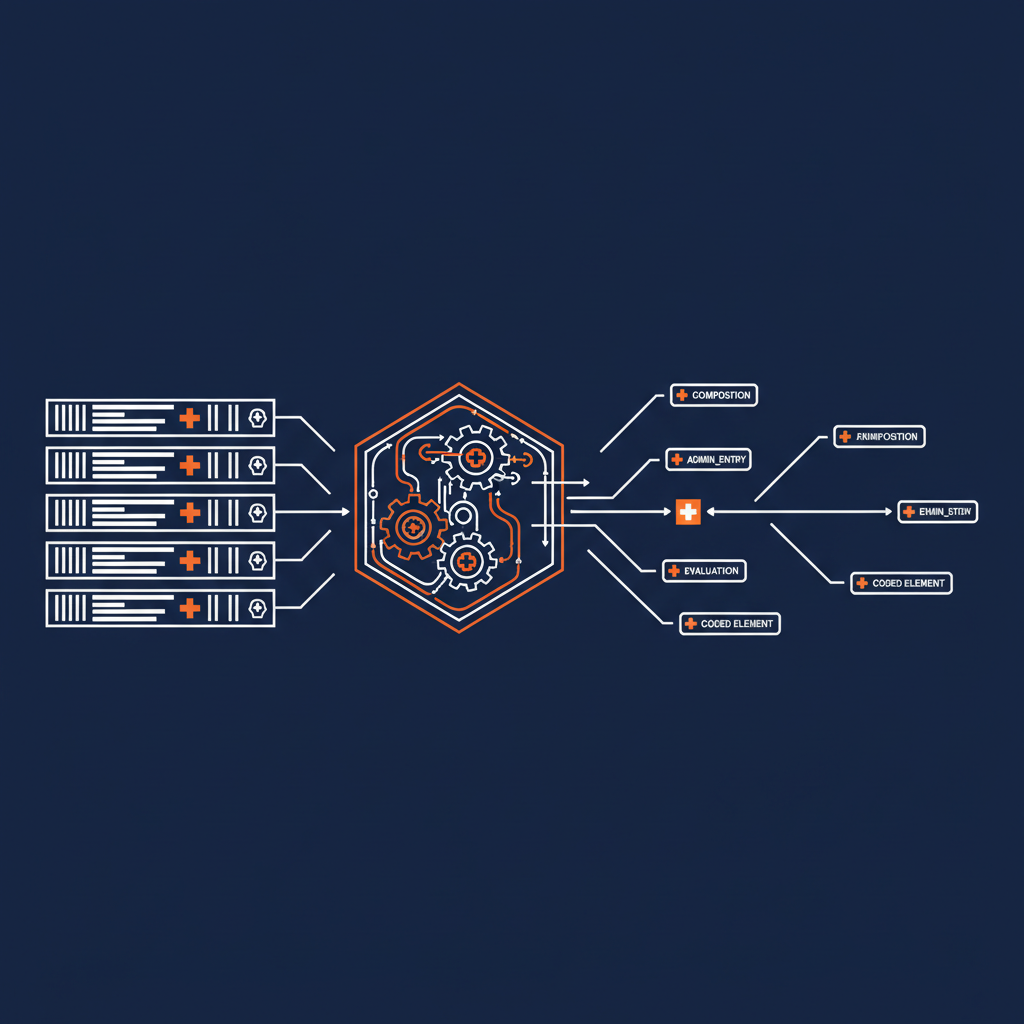
What Gets Produced: The OpenEHR Composition
The Naftiko Capability transforms that pipe-delimited message into a structured OpenEHR composition that any EHDS-compliant system can consume:
{
"_type": "COMPOSITION",
"archetype_node_id": "openEHR-EHR-COMPOSITION.encounter.v1",
"name": { "value": "Inpatient Admission" },
"language": {
"terminology_id": { "value": "ISO_639-1" },
"code_string": "en"
},
"territory": {
"terminology_id": { "value": "ISO_3166-1" },
"code_string": "DE"
},
"composer": {
"_type": "PARTY_IDENTIFIED",
"name": "Dr. Anna Lindqvist",
"external_ref": {
"id": { "_type": "GENERIC_ID", "value": "PROV-2321000016-A123", "scheme": "PROVIDER-ID" },
"namespace": "2.16.840.1.113883.4.1",
"type": "PERSON"
}
},
"context": {
"start_time": { "value": "2024-03-15T08:23:00+01:00" },
"setting": {
"value": "emergency care",
"defining_code": {
"terminology_id": { "value": "openehr" },
"code_string": "227"
}
}
},
"content": [
{
"_type": "ADMIN_ENTRY",
"archetype_node_id": "openEHR-EHR-ADMIN_ENTRY.admission.v0",
"name": { "value": "Admission" },
"subject": {
"_type": "PARTY_IDENTIFIED",
"external_ref": {
"id": { "_type": "HIER_OBJECT_ID", "value": "7812150423" },
"namespace": "2.16.840.1.113883.4.1",
"type": "PERSON"
}
},
"data": {
"_type": "ITEM_TREE",
"items": [
{
"_type": "ELEMENT",
"name": { "value": "Date/time of admission" },
"value": { "_type": "DV_DATE_TIME", "value": "2024-03-15T08:23:00+01:00" }
},
{
"_type": "ELEMENT",
"name": { "value": "Admission type" },
"value": {
"_type": "DV_CODED_TEXT",
"value": "Emergency",
"defining_code": {
"terminology_id": { "value": "openehr" },
"code_string": "EMERGENCY"
}
}
},
{
"_type": "ELEMENT",
"name": { "value": "Ward" },
"value": { "_type": "DV_TEXT", "value": "WARD-23" }
},
{
"_type": "ELEMENT",
"name": { "value": "Room" },
"value": { "_type": "DV_TEXT", "value": "Room-4" }
},
{
"_type": "ELEMENT",
"name": { "value": "Bed" },
"value": { "_type": "DV_TEXT", "value": "Bed-2" }
}
]
}
},
{
"_type": "EVALUATION",
"archetype_node_id": "openEHR-EHR-EVALUATION.problem_diagnosis.v1",
"name": { "value": "Admitting Diagnosis" },
"data": {
"_type": "ITEM_TREE",
"items": [
{
"_type": "ELEMENT",
"name": { "value": "Problem/Diagnosis name" },
"value": {
"_type": "DV_CODED_TEXT",
"value": "Pneumonia, unspecified",
"defining_code": {
"terminology_id": { "value": "ICD-10" },
"code_string": "J18.9"
}
}
}
]
}
}
]
}
The patient’s national identifier is preserved with its correct OID namespace. The provider ID on the attending doctor maps to PARTY_IDENTIFIED. The ICD-10 diagnosis code binds to the proper terminology. The ward, room, and bed from PV1-3 land in discrete, queryable elements. This is not a lossy conversion – it is a structured, governed transformation that retains clinical fidelity.
Patient Discharge: ADT^A03 to OpenEHR Discharge Summary
The discharge is the other bookend. An ADT^A03 fires when a patient leaves the hospital, carrying the final diagnosis, discharge disposition, and critically, the discharge instructions that follow the patient home.
The Capability
naftiko: "1.0.0-alpha1"
info:
label: HL7 v2 Pipe ADT^A03 Patient Discharge to OpenEHR
description: >-
Transforms raw pipe-delimited HL7 v2 ADT^A03 Patient Discharge messages into
OpenEHR-compatible discharge summary compositions with admin_entry for discharge
details, problem_diagnosis for final diagnosis, and clinical_synopsis for
discharge instructions.
tags:
- hl7v2
- openehr
- adt-a03
- patient-discharge
- healthcare
capability:
consumes:
- namespace: hl7-gateway
type: http
baseUri: "https://hl7-gateway.hospital.example.eu/api/v1"
description: >-
Hospital HL7 v2 message gateway. Returns raw pipe-delimited
HL7 v2 ADT^A03 Patient Discharge messages as text/plain.
authentication:
type: bearer
token: "{{HL7_GATEWAY_TOKEN}}"
resources:
- name: discharge-events
path: "/adt/discharges"
operations:
- name: get-discharge-event
method: GET
- namespace: openehr-repo
type: http
baseUri: "https://ehrbase.hospital.example.eu/rest/openehr/v1"
description: >-
OpenEHR clinical data repository (EHRbase). Receives transformed
discharge summary compositions.
authentication:
type: bearer
token: "{{EHRBASE_TOKEN}}"
resources:
- name: compositions
path: "/ehr/{{ehrId}}/composition"
operations:
- name: create-composition
method: POST
exposes:
- type: mcp
port: 3011
namespace: hl7-discharge-tools
description: >-
MCP tools for transforming HL7 v2 ADT^A03 Patient Discharge messages
into OpenEHR discharge summary compositions.
tools:
- name: transform-adt-a03
description: >-
Fetches an ADT^A03 Patient Discharge message, transforms it into an
OpenEHR discharge summary with discharge details, final diagnosis,
and clinical synopsis, then posts it to the OpenEHR repository.
inputParameters:
- name: messageId
type: string
required: true
description: "HL7 message control ID (MSH-10)"
- name: ehrId
type: string
required: true
description: "Target OpenEHR EHR identifier"
steps:
- name: fetch-discharge
type: call
call: hl7-gateway.get-discharge-event
with:
messageId: hl7-discharge-tools.messageId
- name: post-composition
type: call
call: openehr-repo.create-composition
with:
ehrId: hl7-discharge-tools.ehrId
Same pattern: consume, transform, expose. The difference is what gets produced on the other side.
What Gets Consumed: The HL7 v2 Discharge Message
Seven days after admission, the patient is discharged. The ADT^A03 carries the full stay timeline, the final diagnosis (now confirmed, type F instead of admitting type A), and the discharge instructions in the PV2 segment:
MSH|^~\&|EMR_SYSTEM|CENTRAL_HOSPITAL|NATIONAL_REGISTRY|HEALTH_AUTHORITY|20240322143000||ADT^A03^ADT_A03|MSG00042|P|2.5|||AL|NE||UTF-8
EVN|A03|20240322143000|||^Lindqvist^Anna^^^Dr.
PID|1||7812150423^^^NATIONAL_ID&2.16.840.1.113883.4.1&ISO~10045^^^EMR&LOCAL||Mueller^Erik^Johan||19781215|M|||Hauptstrasse 12^^Berlin^^10115^DE^H||+4930123456|||M
PV1|1|I|WARD-23^Room-4^Bed-2|E|||^Lindqvist^Anna^^^Dr.|^Bergman^Lars^^^Dr.||MED|||||||^Lindqvist^Anna^^^Dr.|INP|V201500123456||||||||||||||||01||||||A|||20240315082300|||20240322143000
PV2||||||||||||||||||||Continue antibiotics (Amoxicillin 500mg x3) for 7 days. Follow-up at primary care within 2 weeks. Contact emergency department if shortness of breath or fever above 39C.
DG1|1||J18.9^Pneumonia, unspecified^ICD10|Pneumonia|20240315|F
PV1-36 carries the discharge disposition (01 – discharged to home). PV1-44 and PV1-45 bracket the stay: admitted March 15, discharged March 22. The PV2 segment carries the actual discharge instructions – medication continuation, follow-up appointment, and emergency criteria.
What Gets Produced: The OpenEHR Discharge Summary
The output is a proper discharge summary composition with three content blocks: the administrative discharge record, the confirmed final diagnosis, and the clinical synopsis preserving the discharge instructions:
{
"_type": "COMPOSITION",
"archetype_node_id": "openEHR-EHR-COMPOSITION.discharge_summary.v1",
"name": { "value": "Discharge Summary" },
"language": {
"terminology_id": { "value": "ISO_639-1" },
"code_string": "en"
},
"territory": {
"terminology_id": { "value": "ISO_3166-1" },
"code_string": "DE"
},
"context": {
"start_time": { "value": "2024-03-15T08:23:00+01:00" },
"end_time": { "value": "2024-03-22T14:30:00+01:00" },
"setting": {
"value": "hospital",
"defining_code": {
"terminology_id": { "value": "openehr" },
"code_string": "225"
}
}
},
"content": [
{
"_type": "ADMIN_ENTRY",
"archetype_node_id": "openEHR-EHR-ADMIN_ENTRY.discharge.v0",
"name": { "value": "Discharge" },
"data": {
"_type": "ITEM_TREE",
"items": [
{
"_type": "ELEMENT",
"name": { "value": "Date/time of admission" },
"value": { "_type": "DV_DATE_TIME", "value": "2024-03-15T08:23:00+01:00" }
},
{
"_type": "ELEMENT",
"name": { "value": "Date/time of discharge" },
"value": { "_type": "DV_DATE_TIME", "value": "2024-03-22T14:30:00+01:00" }
},
{
"_type": "ELEMENT",
"name": { "value": "Discharge disposition" },
"value": {
"_type": "DV_CODED_TEXT",
"value": "Discharged to home",
"defining_code": {
"terminology_id": { "value": "local" },
"code_string": "01"
}
}
}
]
}
},
{
"_type": "EVALUATION",
"archetype_node_id": "openEHR-EHR-EVALUATION.problem_diagnosis.v1",
"name": { "value": "Final Diagnosis" },
"data": {
"_type": "ITEM_TREE",
"items": [
{
"_type": "ELEMENT",
"name": { "value": "Problem/Diagnosis name" },
"value": {
"_type": "DV_CODED_TEXT",
"value": "Pneumonia, unspecified",
"defining_code": {
"terminology_id": { "value": "ICD-10" },
"code_string": "J18.9"
}
}
},
{
"_type": "ELEMENT",
"name": { "value": "Diagnostic certainty" },
"value": {
"_type": "DV_CODED_TEXT",
"value": "Confirmed",
"defining_code": {
"terminology_id": { "value": "local" },
"code_string": "at0076"
}
}
}
]
}
},
{
"_type": "EVALUATION",
"archetype_node_id": "openEHR-EHR-EVALUATION.clinical_synopsis.v1",
"name": { "value": "Discharge Instructions" },
"data": {
"_type": "ITEM_TREE",
"items": [
{
"_type": "ELEMENT",
"name": { "value": "Synopsis" },
"value": {
"_type": "DV_TEXT",
"value": "Continue antibiotics (Amoxicillin 500mg x3) for 7 days. Follow-up at primary care within 2 weeks. Contact emergency department if shortness of breath or fever above 39C."
}
}
]
}
}
]
}
The discharge instructions land in a proper clinical_synopsis.v1 archetype. They are no longer buried in a PV2 segment that only HL7-literate systems can parse. Any OpenEHR-connected application, any FHIR bridge, any AI agent with access to the patient’s EHR can now surface these instructions.
From Individual Capabilities to Governed Integration
These capabilities can be executed using the open-source Naftiko Framework in a Docker container. You can aggregate capabilities, introduce multiple steps, and compose different scenarios that produce the standardized and interoperable primary and secondary use outcomes any healthcare provider should be able to deliver in 2026.
The pattern scales. ADT^A01 and ADT^A03 are the admission and discharge bookends, but the same approach applies to ORM^O01 lab orders, ORU^R01 lab results, RDE^O11 medication orders, and every other HL7 v2 message type flowing through European hospital systems. Each becomes a governed capability: declared in YAML, versioned, discoverable, and executable.
What Comes Next
Naftiko is just getting started. An alpha version of the framework is available today, and we are looking for design partners to help us ensure the framework and capabilities work across classic healthcare formats like HL7 v2, as well as modern standards like OpenEHR and FHIR. Our goal is to make sure you can consume and transform whatever you need to confidently integrate AI into healthcare operations.
Consuming legacy data and APIs, and producing the HTTP APIs, MCP tools, and Agent Skills needed to integrate AI into healthcare and clinical environments is just the start. In Q2 2026 we are focused on delivering the Naftiko Fleet alongside the Naftiko Framework, taking all of the capabilities developed by our design partners and ensuring they are:
- Discoverable – Easy to find and make visible as part of business and engineering operations.
- Standardized – Following the standards required as part of any industry regulation in place.
- Compliant – Aligning with government regulations that apply to healthcare integrations.
- Governed – Ensuring the technical and business details of integrations are always managed.
The Naftiko Fleet is where we help healthcare providers connect the dots across all of the standardized integrations put in place, making sure existing investments in patient data can be transformed into what is needed for modern desktop, web, mobile, and AI applications. The Naftiko Framework is how you define what a healthcare provider is capable of. The Naftiko Fleet is how you ensure you can move in whatever direction is needed.
Looking for Design Partners
We are actively looking for healthcare design partners who need to modernize and integrate their existing patient data securely and confidently into AI workflows. The Naftiko Framework and Capabilities specification are both released under the Apache 2.0 license, with the Naftiko Fleet leveraging a commercial open-source approach to help fund what is needed to shape the next generation of healthcare that is interoperable and patient-driven across applications and borders.
If you are working with HL7 v2, OpenEHR, FHIR, or any combination of healthcare data standards and want to explore how governed capabilities can accelerate your integration roadmap, we want to hear from you.